1. Diverse photoreceptors in plants
Many civilizations, including the sun god of ancient Egypt, thought that the blessings of sunlight were the source of life. In fact, the survival of all life, including humans, is supported by the photosynthesis of plants that capture solar energy. Plants that perform photosynthesis have no means of transportation except for some algae.Therefore, it is necessary to monitor various changes in the external environment and respond appropriately to the place to survive. Among various environmental information, light is especially important information for plants that perform photosynthesis.
In the process of evolution, plants acquired phytochrome , which mainly receives light in the red light region, and multiple blue light receptors , including his lyptochrome and phototropin, in order to sense the light environment. .. In addition to these, an ultraviolet light receptor named UVR8 was recently discovered. In this paper, we explain the latest image of the molecular structure and function of these various plant photoreceptors (Fig. 1), focusing on phytochrome and phototropin.
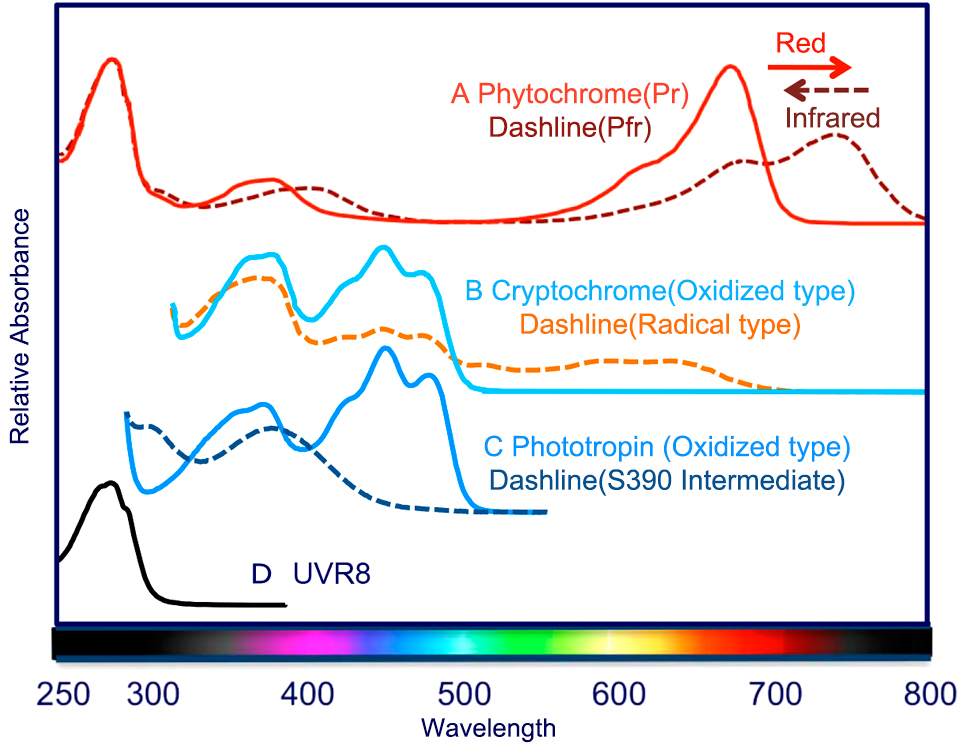
Figure 1
Ultraviolet-visible absorption spectra of phytochrome, cryptochrome, phototropin, and UVR8. The dashed line represents each bioactive absorption spectrum.
2. Phytochrome; red-far red photoreversible molecular switch
What is phytochrome?
Phytochrome is a photochromic photoreceptor, and has two absorption types, a red light absorption type Pr (absorption maximum wavelength of about 665 nm) and a far-red light absorption type Pfr (730 nm). Reversible light conversion between the two by red light and far-red light, respectively(Fig. 1A, solid line and broken line). In general, Pfr is the active form that causes a physiological response. With some exceptions, phytochrome can be said to function as a photoreversible molecular switch. The background of the discovery is as follows. There are some types of plants that require light for germination (light seed germination). From that study, it was found that germination was induced by red light, the effect was inhibited by subsequent far-red light irradiation, and this could be repeated, and the existence of photoreceptors that reversibly photoconvert was predicted. In 1959, its existence was confirmed by the absorption spectrum measurement of the yellow sprout tissue, and it was named phytochrome. Why does the plant have a sensor to distinguish between such red light and far-red light? There is no big difference between the red and far-red light regions in the open-field spectrum of sunlight, but the proportion of red light is greatly reduced due to the absorption of chloroplasts in the shade of plants. Similar changes in light quality occur in the evening sunlight. Plants perceive this difference in light quality as the ratio of Pr and Pfr, recognize the light environment, and respond to it.
Subsequent studies have revealed that it is responsible for various photomorphogenic reactions such as photoperiodic flowering induction, shade repellent, and deyellowing (greening). Furthermore, with the introduction of the model plant Arabidopsis thaliana (At) and the development of molecular biological analysis methods, research has progressed dramatically, and his five types of phytochromes (phyA-E) are present in Arabidopsis thaliana. all right. With the progress of the genome project, Fi's tochrome-like photoreceptors were found in cyanobacteria, a photosynthetic prokaryotes other than plants. Furthermore, in non-photosynthetic bacteria, a homologue molecule called bacteriophytochrome photoreceptor (BphP) was found in Pseudomonas aeruginosa (Pa) and radiation-resistant bacteria (Deinococcus radiodurans, Dr).
Domain structure of phytochrome molecule
Phytochrome molecule can be roughly divided into N-terminal side and C-terminal side region. PAS (Per / Arndt / Sim: blue), GAF (cGMP phosphodiesterase / adenylyl cyclase / FhlA: green), PHY (phyto-chrome: purple) 3 in the N-terminal region of plant phytochrome (Fig. 2A) There are two domains and an N-terminal extension region (NTE: dark blue), and phytochromobilin (PΦB), which is one of the ring-opening tetrapyrroles, is thioether-bonded to the system stored in GAF as a chromophore. ing. PAS is a domain involved in the interaction between signal transduction-related proteins, and PHY is a phytochrome-specific domain. There are two PASs and her histidine kinase-related (HKR) domain (red) in the C-terminal region, but the histidine essential for kinase activity is not conserved.
On the other hand, in Cph1 (Fig. 2B) of the cyanobacteria Synechocystis (Syn), which has an N-terminal region similar to plant phytochrome, phycocyanobilin, which has a shorter π electron-conjugated system than PΦB, binds to GAF cysteine. The C-terminus is histidine kinase (HK: red). Furthermore, in BphP, the chromophore is biliverdin, and it is bound to cysteine present upstream of the N-terminal of PAS instead of GAF (Fig. 2C). Despite the different chromophores and their binding sites, both Cph1 and BphP show reversible photoconversion reactions between Pr and Pfr, much like plant phytochromes.
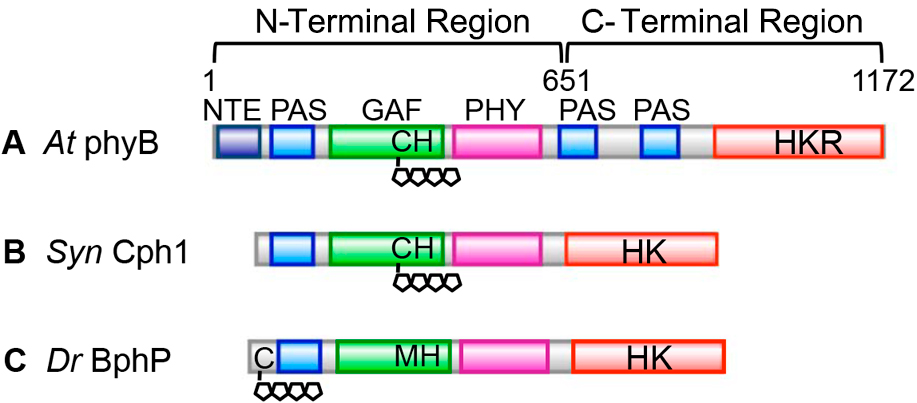
Figure 2
Domain structure of phytochrome. (A) One of the plant phytochromes AtphyB, (B) Cyanobacterial phytochrome SynCph1, (C) One of the bacteriophytochromes DrBphP. The four pentagons represent the tetrapyrrole chromophore and C represents the bound cysteine. See the text for details.
Photoreaction with photoisomerization and proton transfer of phytochrome chromophore
Fig. 3A shows the molecular structure of PΦB, and B shows the photoreaction. By photoexcitation, Pr undergoes photoisomerization and subsequent proton transfer reaction, similar to other π-electron-conjugated double bond rhodopsin and bacteriorhodopsin chromophore retinal. In isomerization, a reaction from 15Za to 15Ea occurs between the C ring and the D ring, followed by a transfer reaction of the C ring nitrogen atom-bonded proton. Also, the photochemical pathway from Pfr to Pr in the reverse reaction is different from the pathway from Pr to Pfr. Furthermore, apart from the photoreaction, a thermal dark regression reaction from Pfr to Pr also occurs. Although the reversibility of the photochemical reaction can be seen only in GAF, PAS and PHY are required to maintain the normal absorption spectra of Pr and Pfr. In addition, PHY and NTE play important roles in the stability of Pfr, and the loss of these plays a faster dark regression reaction from Pfr to Pr.
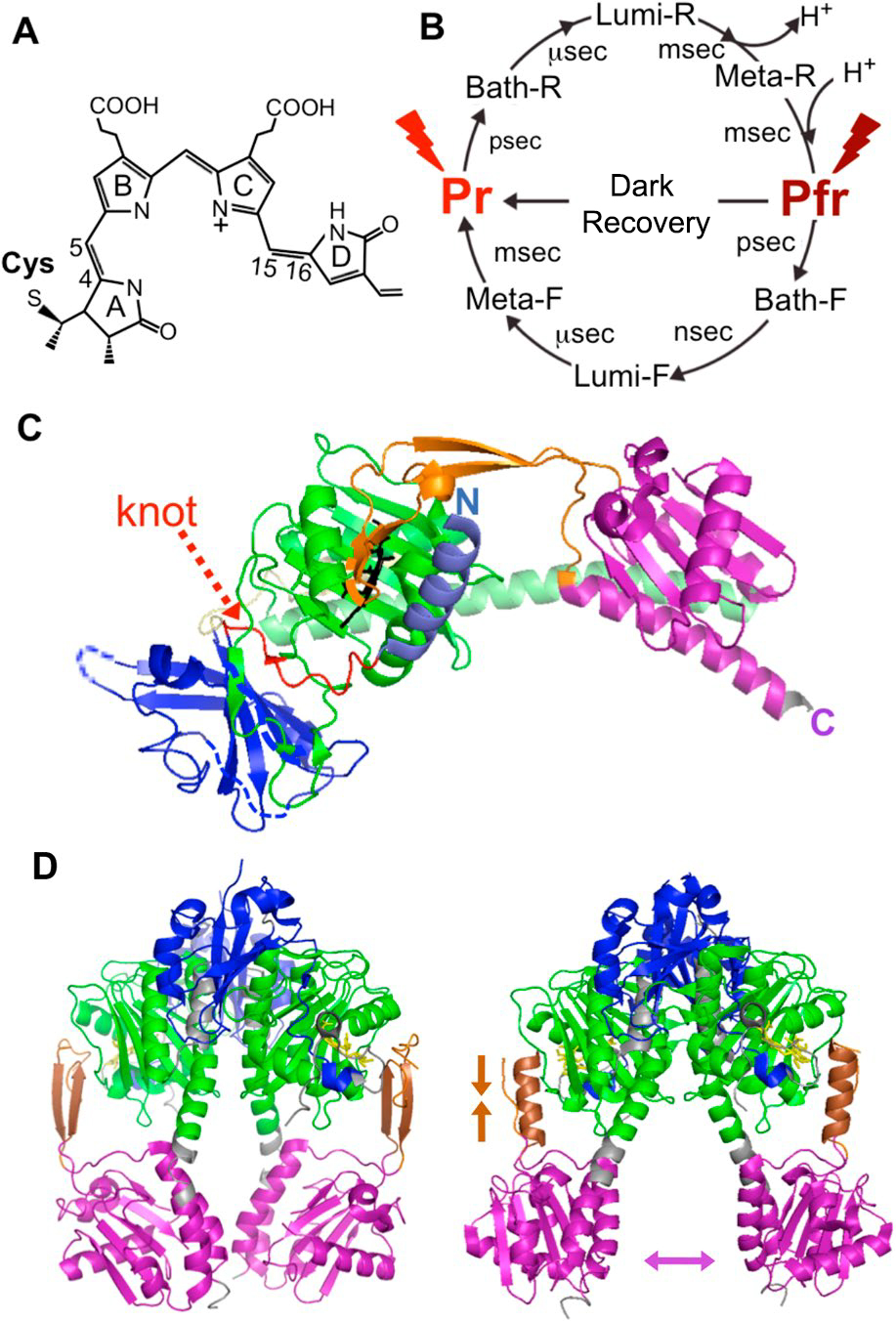
Figure 3
(A) Molecular structure of Pr type PΦB. (B) Phytochrome photoconversion reaction. (C) Crystal structure of Cph1 N-terminal region sample. (D) DrBphP N-Three-dimensional structure of the terminal region under dark (Pr, left) and under bright (Pfr, right). The colors of each domain in (C) and (D) correspond to Fig. 2. See the text for details.
Phytochrome crystal structure with knots
In 2005, the first crystal structure was solved for Pr in the N-terminal region lacking the PHY of DrBphP, and then the crystal structure in the N-terminal region containing PHY was Cph1 (Fig. 3C) and It was announced at PaBphP and others. Interestingly, despite the different binding positions of the chromophores in BphP and Cph1, the loops formed by the looped inserted amino acid sequences conserved in GAF (green) are N in PAS (blue). -Several tens of amino acid residues (red) in the upstream terminal region passed through to form a knot (arrow in Fig. 3C). This light-sensing knot forms part of the chromophore (black bar model) coupling pocket and is thought to be deeply involved in the light conversion reaction. The force required to solve the knot is 73 pN, which is not large enough to stabilize the protein structure, according to studies using an atomic force microscope. From the Magic Angle Spinning NMR measurement results, it has been reported that the environment around the chromophore changes from a "soft" state to a "hard" state with the photoconversion of Pr to Pfr, and knot is the photoisomerization of the chromophore. It is thought that it plays a role in correctly guiding the protein structural change in order to carry out the above accurately. The GAF is a long alpha helix (light green) called the connecting helix, which he is connected to the PHY (purple). From the PHY, a structure (orange) called tongue extends toward the GAF chromophore pocket, which is also involved in the spectroscopic properties of the phytochrome and plays a major role in structural changes associated with photoconversion.
Molecular structural changes associated with the photoconversion reaction of bacteriophage phytochrome
What kind of structural changes are caused in the protein part due to the reaction of the chromophore? Last year, the dark state (Fig. 3D left) and light state (same right) structures of DrBphP were reported based on his small-angle X-ray scattering (SAXS) and crystal structure. According to this, DrBphP exists as a parallel dimer, and the middle part (brown part) of the tongue extending from PHY to GAF forms an antiparallel β sheet consisting of two β strands in the dark state. , In the clear state, it is a single α-helix. As a result, the length of the tongue is shortened by 2.5 Å and the PHY is attracted to the GAF, and he is a dimer with the PHY wide open in a Y shape. These changes reflect the structural changes associated with the Pr → Pfr photoconversion reaction, and are thought to be related to the activity control of HK, which is the signature module on the C-terminal side. The Pr dimer image of the full-length DrBphP obtained from the single molecule measurement by cryo-EM shows a structure in which the HK region existing at the tip of the PHY is in contact and twisted, and this structure is formed by photoreaction. I am interested in how it changes.
Function and structure of N- and C-terminal region of plant phytochrome
The N-terminal region of plant phytochrome bound with PΦB shows reversible photoconversion and functions as an optical sensory module. Interestingly, in Arabidopsis phyB, the optical sensory module has been shown to have a signaling function in addition to a photoreceptive function. Pr localized in the cytoplasm translocates to the nucleus when photoconverted to Pfr. In the nucleus, Pfr binds to gene transcription factors such as PIF (phytochrome-interacting factor), which is a bHLH-type gene transcription factor, and induces their degradation by the ubiquitin-proteasome system. PIF functions as a suppressor or promoter of protein gene transcription involved in various photomorphogenesis reactions, and the degradation or promotion of PIF is released by degradation of PIF, leading to photomorphogenesis. Functional complementation experiments of the white-spotted phyB-deficient strain showed that the optical sensory module of phyB has a signaling function. On the other hand, the C-terminal region has a nuclear localization signal and a dimerization site and is considered to function as a control module. With exceptions, plant phytochromes generally function as homodimers. It is considered that structural changes such that the nuclear localization signal is exposed occur in the control module due to the optical conversion.
Last year, the Pr crystal structure of the Arabidopsis phyB optical sensory module (lacking 90 amino acids at the N-terminal and 27 amino acids at the C-terminal) was reported. The crystal structure is a parallel dimer similar to DrBphP, and has a knot or β-sheet type tongue structure. However, in the Pr of DrBphP, the PHYs of both subunits warp in an arc shape and show an O-leg-shaped structure (Fig. 3D left), whereas in the Pr of phyB, the PHY extends and contacts in a II shape. This difference in molecular structure may reflect the difference in the signal transduction mechanism between DrBphP and phyB, the former binding to HK activity regulation and the latter binding to PIF and the like. The SAXS-based dimer structure of phyA, which lacks his NTE in Alaskan peas, is quite different from the above-mentioned full-length DrBphP dimer electron microscope image, which also reflects the difference in function between the two. May. Although it may be difficult to prepare a full-length crystal, it is awaited to elucidate the crystal structure of Pr and Pfr of the full length of the photosensory module of the plant phy in order to elucidate the photoreceptive molecular mechanism of the plant.
3. Phototropin; photosynthetic efficiency optimized blue light receptor
What is phototropin?
Charles Darwin, who is famous for his theory of evolution, wrote in his book "The power of move-ment in plants" published in 1882 that plants bend toward blue light. Approximately 100 years later, the protein nph1 (nonphoto-tropic hypocotyl 1) encoded by one of the causative genes of Arabidopsis mutants causing phototropic abnormalities was identified as a blue photoreceptor. Later, another isotype npl1 was found and renamed phototropin 1 (phot1) and 2 (phot2), respectively. In addition to phototropism, phototropin is damaged by chloroplast photolocalization (chloroplasts move through the epidermal cells of the leaves and gather on the cell surface under appropriate light intensity for photosynthesis. As a photoreceptor for reactions such as escaping to the side of cells under dangerous strong light) and stomata (reactions that open stomata to optimize the uptake of carbon dioxide, which is the rate-determining process of photosynthetic reactions). It became clear that it worked. In this way, phototropin can be said to be a blue light receptor responsible for optimizing photosynthetic efficiency.
Domain structure and LOV photoreaction of phototropin molecule
Phototropin molecule has two photoreceptive domains (LOV1 and LOV2) called LOV (Light-Oxygen-Voltage sensing) on the N-terminal side, and serine / on the C-terminal side. It is a protein kinase that forms threonine kinase (STK) (Fig. 4Aa) and whose activity is regulated by light. LOV is one molecule as a chromophore, he binds FMN (flavin mononucleotide) non-covalently. The LOV forms an α/βfold, and the FMN is located on a β-sheet consisting of five antiparallel β-strands (Fig. 4B). The FMN in the ground state LOV shows the absorption spectrum of a typical oxidized flavin protein with a triplet oscillation structure and an absorption maximum wavelength of 450 nm, and is called D450 (Fig. 1C and Fig. 4E). After being excited to the singlet excited state by blue light, the FMN shifts to the triplet excited state (L660t *) due to intersystem crossing, and then the C4 (Fig. 4C) of the isoaroxazine ring of the FMN is conserved in the vicinity. It forms a transient accretionary prism with the tain (red part in Fig. 4B Eα) (S390I). When this cysteine is replaced with alanine (C / A substitution), the addition reaction does not occur. The effect of adduct formation propagates to the protein moiety, causing kinase activation (S390II). After that, the formed cysteine-flavin adduct spontaneously dissociates and returns to the original D450 (Fig. 4E, dark regression reaction).
Phototropin kinase activity control mechanism by LOV2
Why does phototropin have two LOVs? Atphot1 was found as a protein that is rapidly autophosphorylated when irradiated with blue light.
The effect of the above C / A substitution on this self-phosphorylation reaction and phototropism was investigated, and LOV2 is the main photomolecular switch in both self-phosphorylation and phototropism. It turns out that it functions as. After that, from experiments using artificial substrates, STK has a constitutive activity, LOV2 functions as an inhibitory domain of this activity, and the inhibition is eliminated by photoreaction, while LOV1 is kinase light. It was shown to modify the photosensitivity of the activation reaction. In addition to this, LOV1 was found to act as a dimerization site from the crystal structure and his SAXS. What kind of molecular mechanism does LOV2 use to photoregulate kinase activity? The following two modules play important roles in this intramolecular signal transduction.
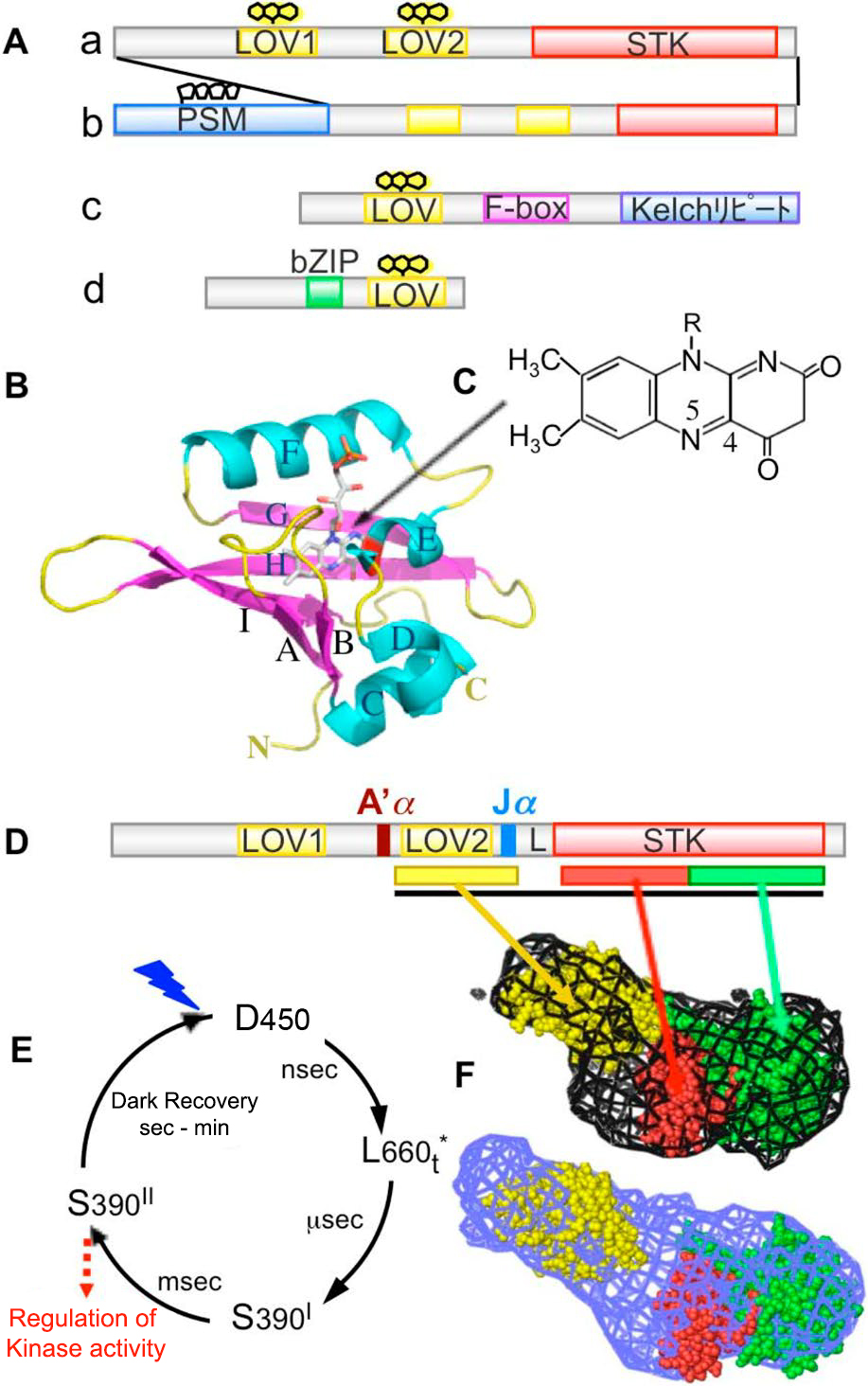
Figure 4
(A) Domain structure of LOV photoreceptors. a: Phototropin b: Neochrome c: FKF1 family protein d: Aureochrome (B) Crystal structure of auto barley phot1 LOV2. (C) Structure of FMN isoaroxazine ring. (D) Schematic diagram of the functional domain and module of Arabidopsis thaliana phot1. L, A'α, and Jα represent linker, A'α helix, and Jα helix, respectively. (E) LOV photoreaction. (F) Molecular structure model (mesh) of the LOV2-STK sample (black line) containing A'α of phot2 obtained based on SAXS under dark (top) and under bright (bottom). The yellow, red, and green space-filled models represent the crystal structures of LOV2-Jα, protein kinase A N-lobe, and C-robe, respectively, and black represents FMN. See the text for details.
1) Jα. LOV2 C of oat phot1-to α immediately after the terminus
Rix (Jα) is present (Fig. 4D), which interacts with the β-sheet (Fig. 4B) that forms the FMN-bound scaffold of LOV2 in the dark, but unfolds and dissociates from the β-sheet with photoreaction. It was shown by NMR that it does. According to the crystal structure of LOV2-Jα, this Jα is located on the back surface of the β sheet and mainly has a hydrophobic interaction. The formation of S390II causes twisting of the isoaroxazine ring and protonation of N5 (Fig. 4C).
As a result, the glutamine side chain present on his Iβ strand (Fig. 4B) in the β-sheet rotates to form a hydrogen bond with this protonated N5. Jα interacts with this his Iβ strand, and these changes are thought to cause the unfold-ing of Jα and dissociation from the β-sheet described above. Experiments such as amino acid substitution of Iβ strands revealed that kinases exhibit constitutive activity when this interaction is eliminated, and that Jα plays an important role in photoactivation of kinases.
2) A'α / Aβ gap.
Recently, several results have been reported showing the involvement of amino acids near the A'α helix (Fig. 4D) located upstream of the N-terminal of LOV2 in kinase photoactivation. Therefore, he investigated the role of this A'α and its neighboring amino acids in kinase photoactivation, photoreaction, and Jα structural change for Atphot1. The LOV2-STK polypeptide (Fig. 4D, underlined in black) was used as a photocontrollable kinase for kinase activity analysis. As a result, it was found that the photoactivation of the kinase was abolished when amino acid substitution was introduced into the A'α / Aβ gap between A'α and Aβ of the LOV2 core. Interestingly, he had no effect on the structural changes in Jα examined on the peptide map due to the photoreaction of LOV2 or trypsin degradation. Therefore, the A'α / Aβ gap is considered to play an important role in intramolecular signal transduction after Jα.
Structural changes detected by SAXS
Structural changes of Jα have been detected by various biophysical methods other than NMR, but structural information on samples including up to STK is reported only by his results to his SAXS. Not. The SAXS measurement of the Atphot2 LOV2-STK polypeptide showed that the radius of inertia increased from 32.4 Å to 34.8 Å, and the molecular model (Fig. 4F) obtained by the ab initio modeling software GASBOR is that of LOV2 and STK. It was shown that the N lobes and C lobes lined up in tandem, and the relative position of LOV2 with respect to STK shifted by about 13 Å under light irradiation. The difference in the molecular model between the two is considered to reflect the structural changes that occur in the Jα and A'α / Aβ gaps mentioned above.
Two phototropins with different photosensitivity
In the phototropic reaction of Arabidopsis Arabidopsis, Arabidopsis responds to a very wide range of light intensities from 10–4 to 102 μmol photon / sec / m2. At that time, phot1 functions as an optical sensor in a wide range from low light to strong light, while phot2 reacts with light stronger than 1 μmol photon / sec / m2. What is the origin of these differences? As is well known, animal photoreceptors have a high photosensitivity due to the abundance of rhodopsin and the presence of biochemical amplification mechanisms. The exact abundance of phot1 and phot2 in vivo is unknown, but interesting results have been obtained in terms of amplification.
The light intensity dependence of the photoactivation of the LOV2-STK polypeptide used in the above kinase analysis was investigated.
It was found that phot1 was about 10 times more photosensitive than phot2. On the other hand, when the photochemical reactions of both were examined, it was found that the rate of the dark return reaction of phot1 was about 10 times slower than that of phot2. This result indicates that the longer the lifetime of S390II, which is in the kinase-activated state, the higher the photosensitivity of kinase activation. This correlation was further confirmed by extending the lifespan of her S390II with amino acid substitutions. This alone cannot explain the widespread differences in photosensitivity between phot1 and phot2, but it may explain some of them. Furthermore, it is necessary to investigate in detail protein modifications such as phosphorylation and the effects of phot interacting factors on photosensitivity.
Other LOV photoreceptors Among fern plants and green algae, phytochrome ɾphotosensory module (PSM) on the N-terminal side and chimera photoreceptor with full-length phototropin on the C-terminal side, neochrome (Fig. There are types with 4Ab). It has been reported that some neochromes play a role in chloroplast photolocalization as a red light receiver. It is considered that fern plants have such a chimera photoreceptor in order to survive in a habitat such as undergrowth in a jungle where only red light reaches.
In addition to this, plants have only one LOV domain, and three proteins involved in the degradation of photomorphogenesis-related proteins, FKF1 (Flavin-binding, Kelch repeat, F-box 1, ZTL (ZEITLUPE)), LKP2 ( There are LOV Kelch Protein2) (Fig. 4Ac) and aureochrome (Fig. 4Ad), which has a bZip domain on the N-terminal side of LOV and functions as a gene transcription factor.
4. Cryptochrome and UVR8
Cryptochrome is one of the blue photoreceptors and forms a superfamily with the DNA photoreceptor photolyase. It has FAD (flavin adenine dinucle-otide) as a chromophore and tetrahydrofolic acid, which is a condensing pigment. The ground state of FAD is considered to be the oxidized type, and the radical type (broken line in Fig. 1B) generated by blue light irradiation is considered to be the signaling state. The radical type also absorbs in the green to orange light region, and may widen the wavelength region of the plant morphogenesis reaction spectrum. Cryptochrome uses blue light to control physiological functions similar to phytochrome.
It was identified as a photoreceptor from one of the causative genes of UVR8 Arabidopsis thaliana, and the chromophore is absorbed in the UVB region by a Trp triad consisting of three tryptophans (Fig. 1D). It is involved in the biosynthesis of flavonoids and anthocyanins that function as UV scavengers in plants.
Conclusion
It is thought that plants have acquired various photoreceptors necessary for their survival during a long evolutionary process. The photoreceptors that cover the existing far-red light to UVB mentioned here are considered to be some of them. More and more diverse photoreceptor genes are conserved in cyanobacteria and marine plankton. By examining these, it is thought that the understanding of plant photoreceptors will be further deepened.